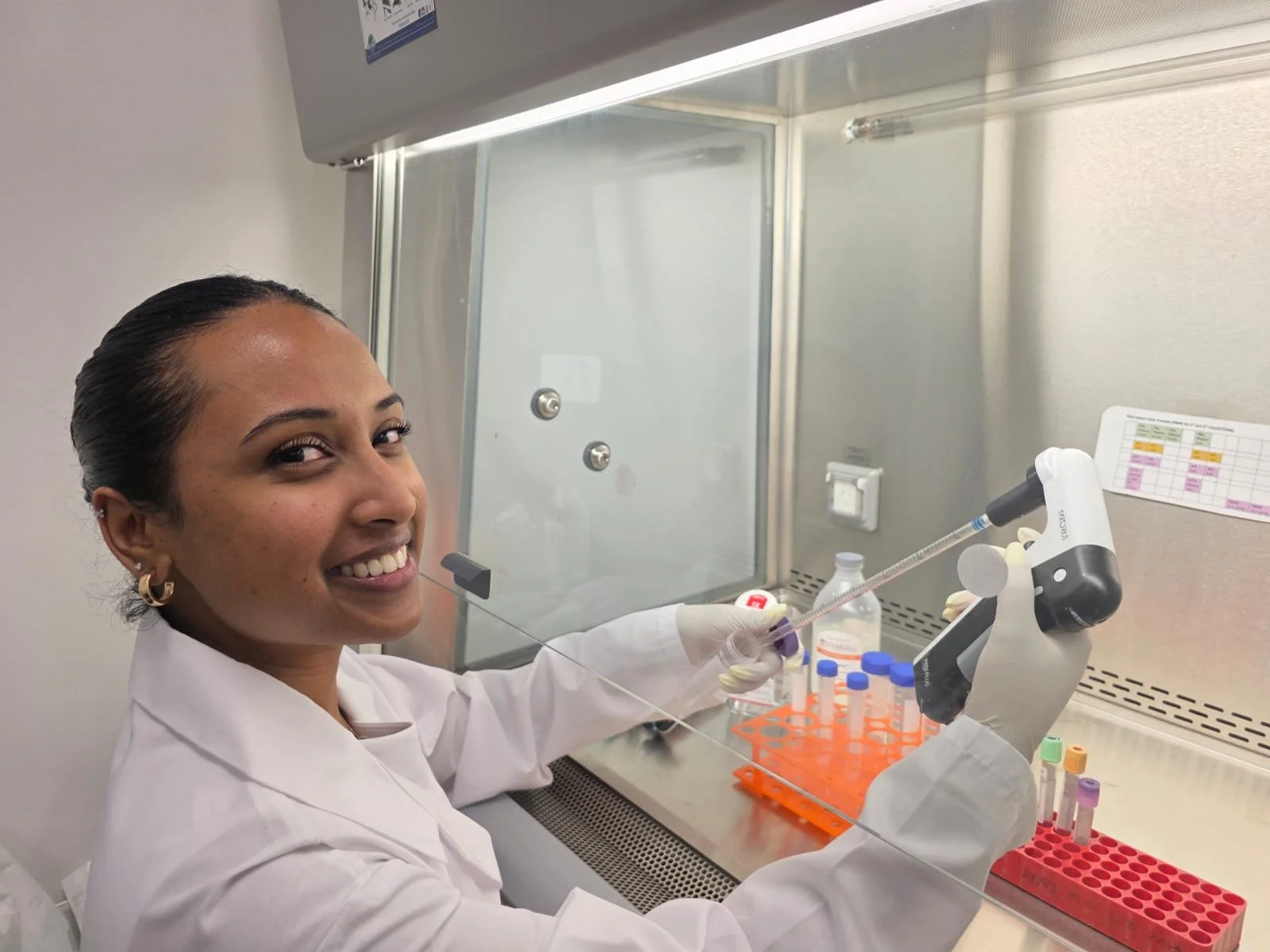
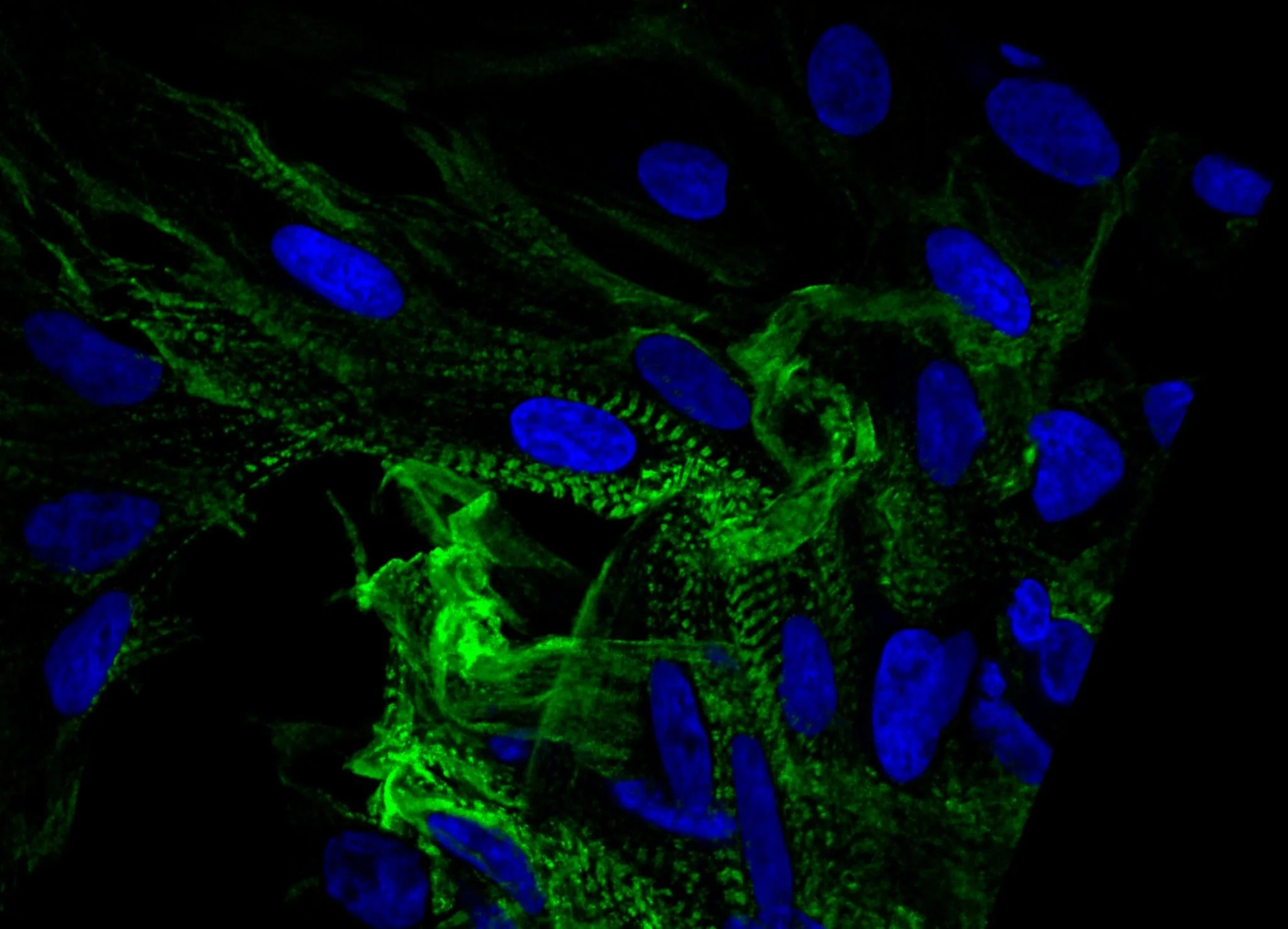
PrecisionGo Capabilities
PrecisionGo offers biomedical and clinical researchers access to state-of-the-art infrastructure, expertise and advanced technologies critical for the development of personalized therapies. Key to this goal is delivering our capabilities through fully integrated end-to-end pipelines to streamline and fast-track biomarker discovery and validation for a wide spectrum of diseases.
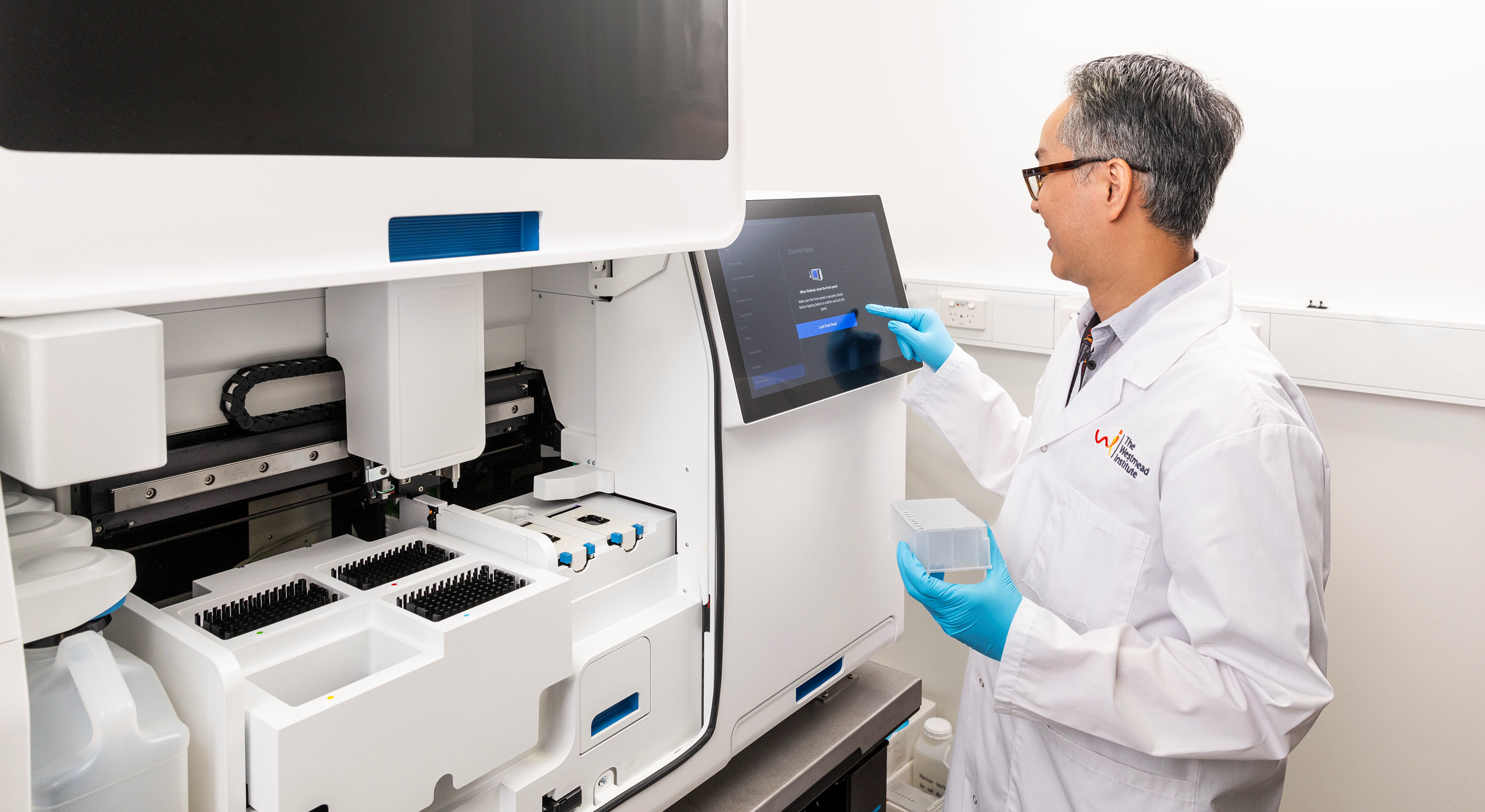

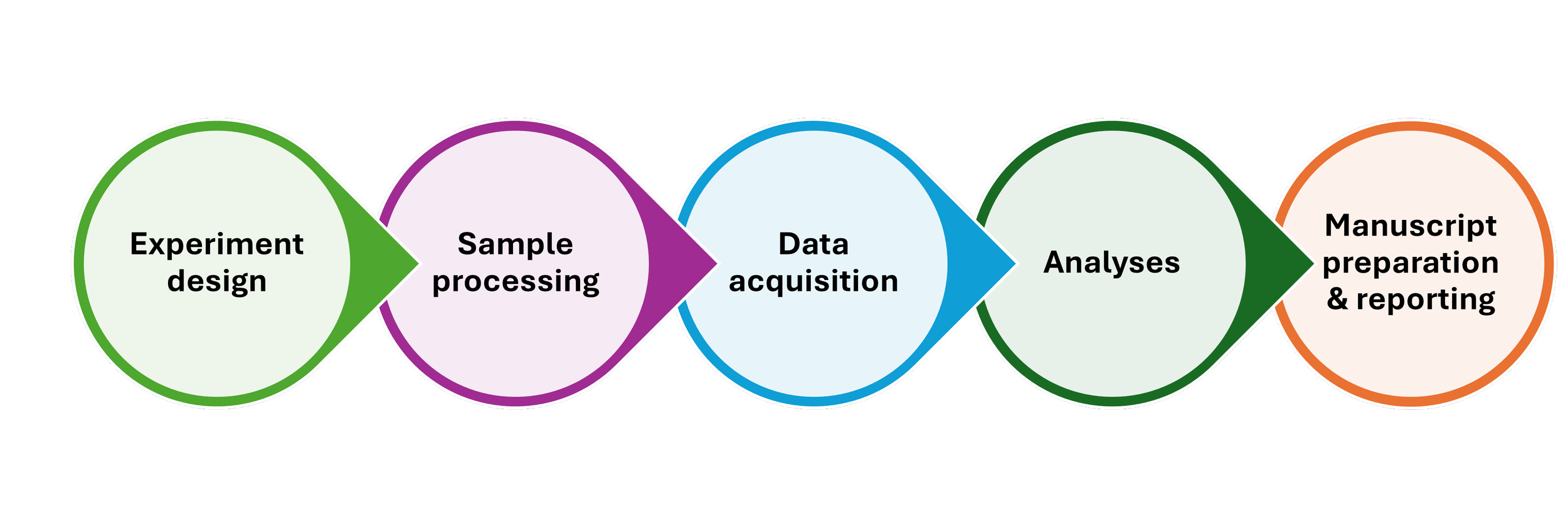
The PrecisionGo Pathway

Our Technologies and Expertise
PrecisionGo is imbedded in the WRH Core Facilities providing world-class equipment and expertise for researchers and industry, offering access to over 100 cutting-edge instruments supported spanning 10 technology field by more than 20 highly skilled technology specialists. We are currently supporting >800 projects from >200 groups from 65 organisations.
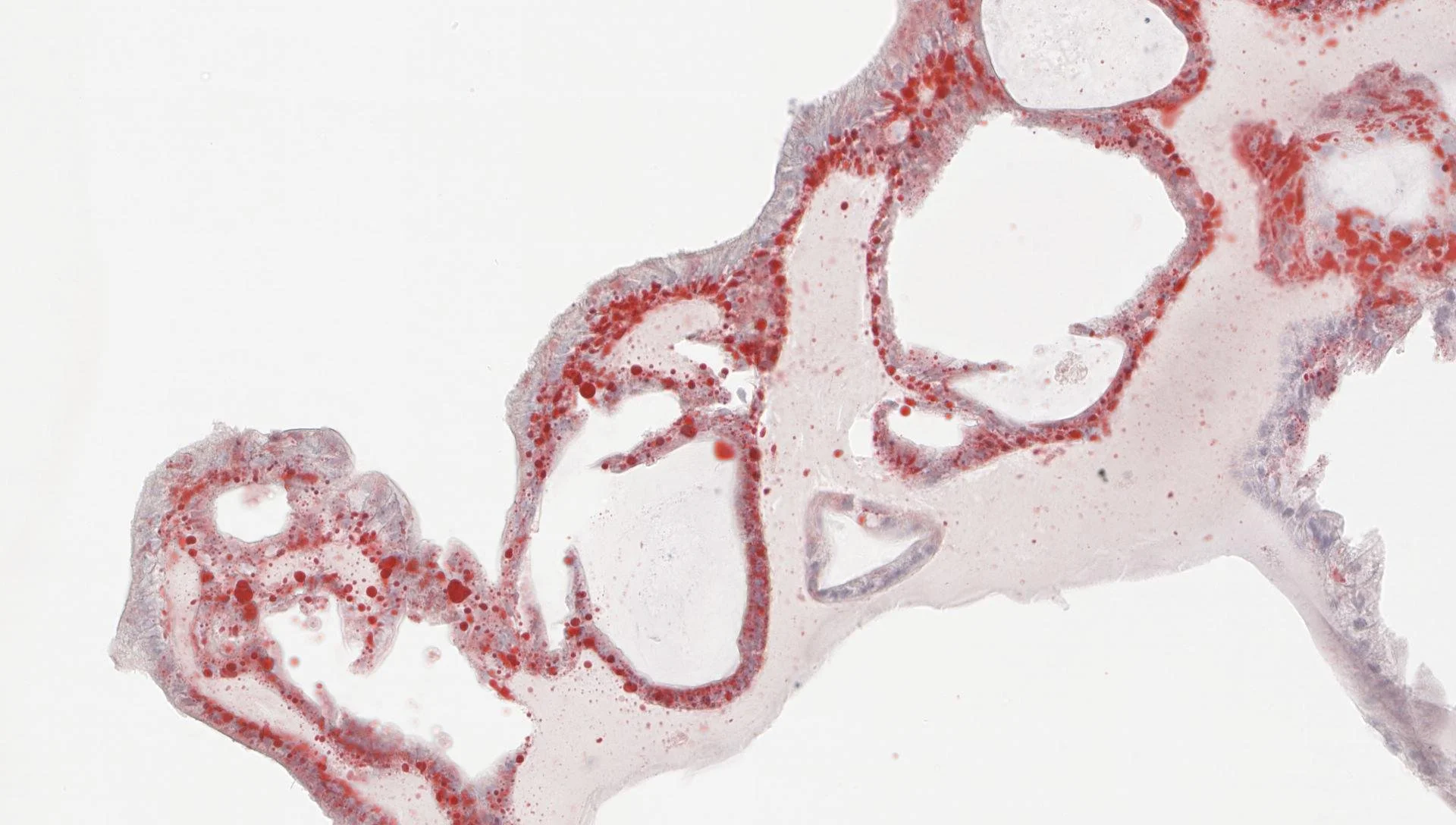
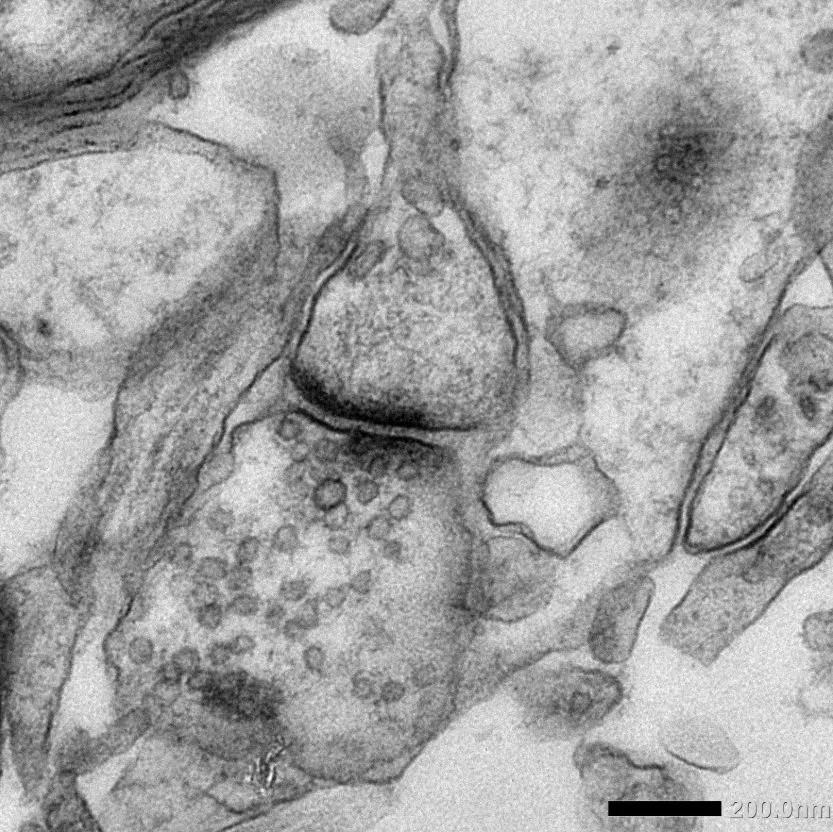
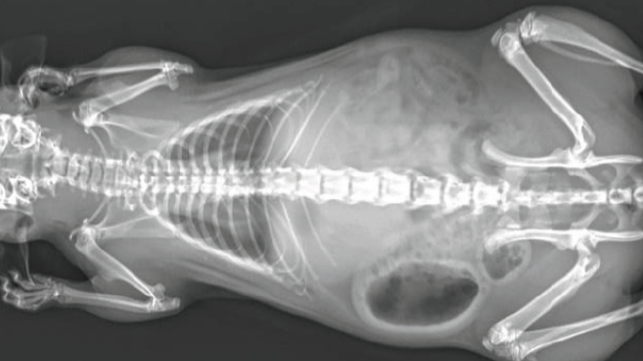
PrecisionGo operates through a single point of contact to manage your project from start to finish
From initial project consultation with our experienced senior specialists to generating data for high-impact publications, PrecisionGo supports a wide range of precision medicine projects for biomarker discovery and validation through to clinical translation. This involves:
Support in aligning study design, analysis, and downstream translation
Coordinating access and utilization across different technology platforms to meet project needs
Enabling secure integration of multiomic data with clinical datasets
Project resource and data management
Aligning governance, quality, and oversight from the outset
Our Pipelines to Accelerate Biomarker Discoveries

Newly Developed Pipelines
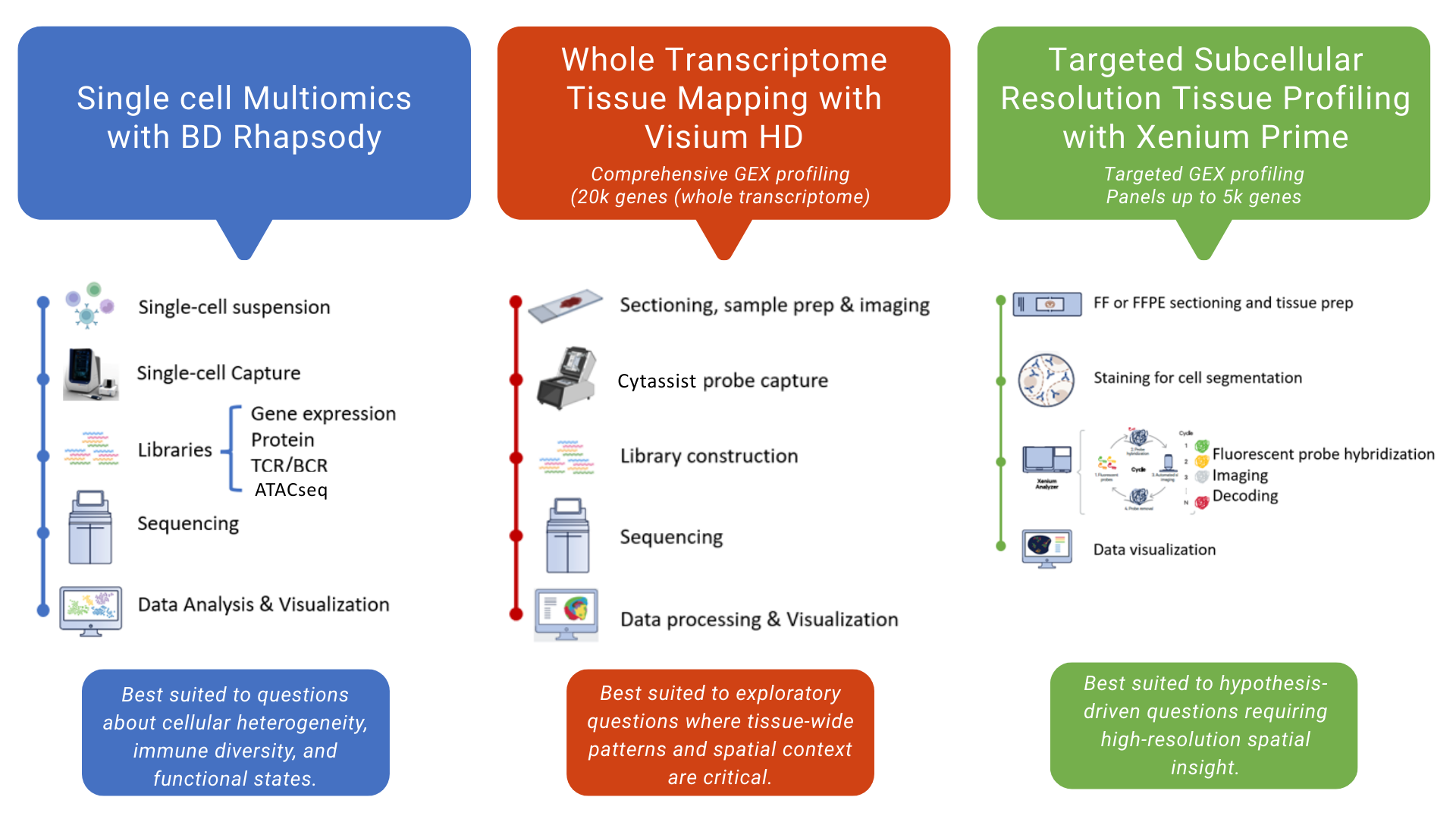
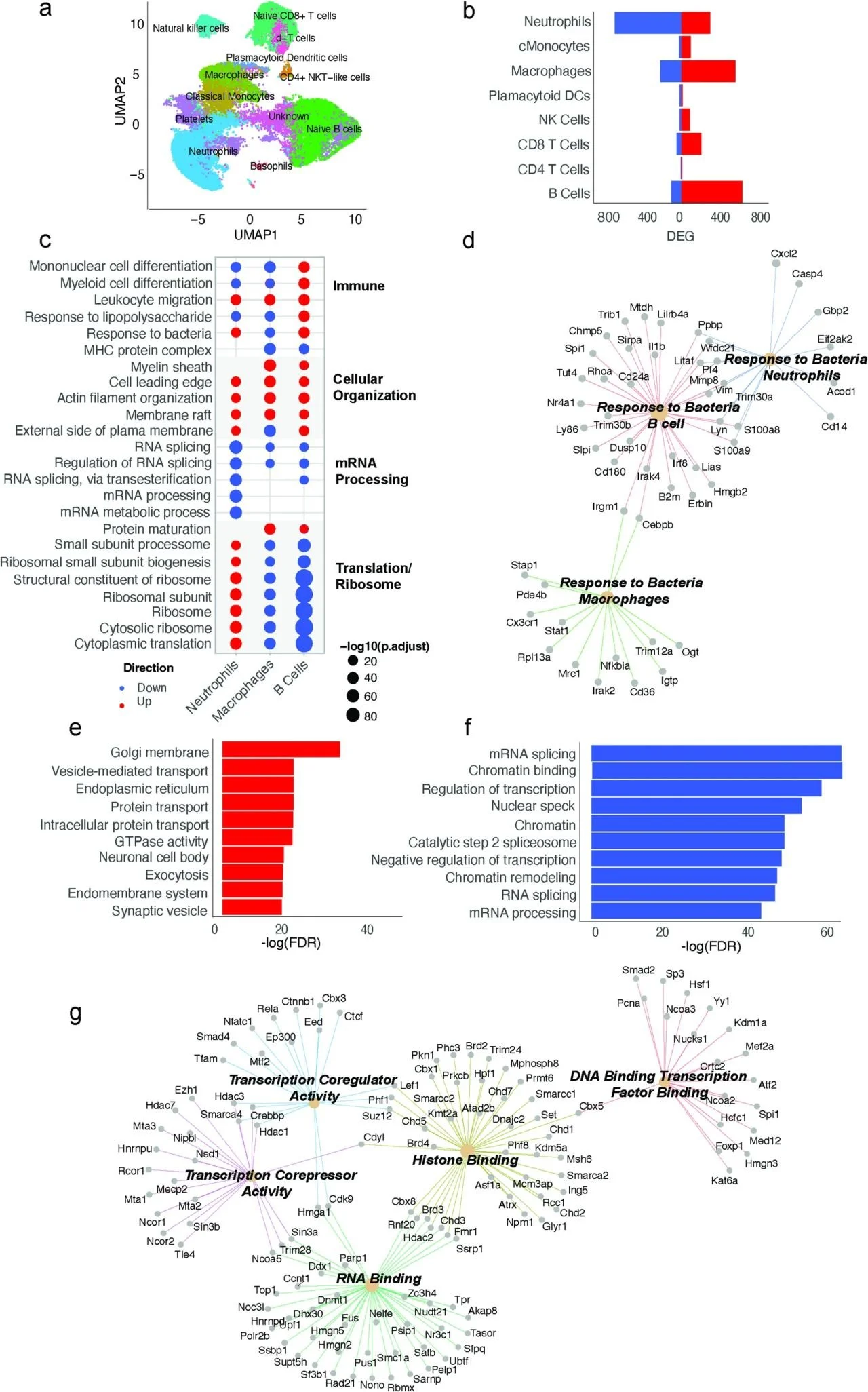
PrecisionGo supports integrated pipelines that are designed to accelerate biomarker discovery within a single, coherent workflow. In addition to traditional genomic profiling pipelines using next generation sequencing, PrecisionGo offers newly developed pipelines for multiomic single cell and spatial transcriptomics. These current generation technologies allow comprehensive and targeted profiling of key cellular and molecular biomarkers to improve disease risk stratification, enable earlier diagnosis and accurate prognosis, predict treatment responses and ultimately improve clinical decisions for better patient outcomes.
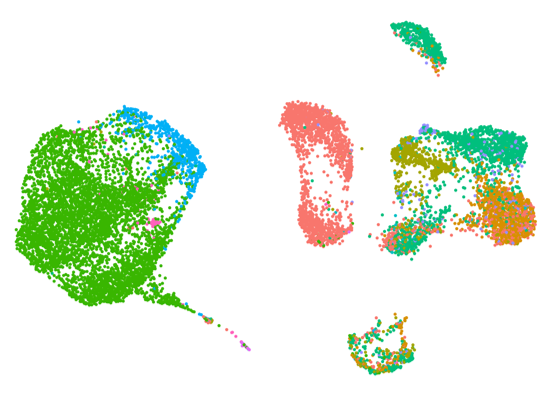
Single-cell multiomic profiling
This pipeline enables integrated analysis of individual cells by combining whole transcriptome analysis with additional molecular and cellular features, such as immune receptor diversity and protein expression. It supports studies seeking to:
characterise cellular heterogeneity
link transcriptional states to immune or functional phenotypes
explore clonal diversity of immune cells and lineage relationships
Through generating multiple data types from the same cells, this pipeline supports richer interpretation and more precise biological insight than single-modality approaches.
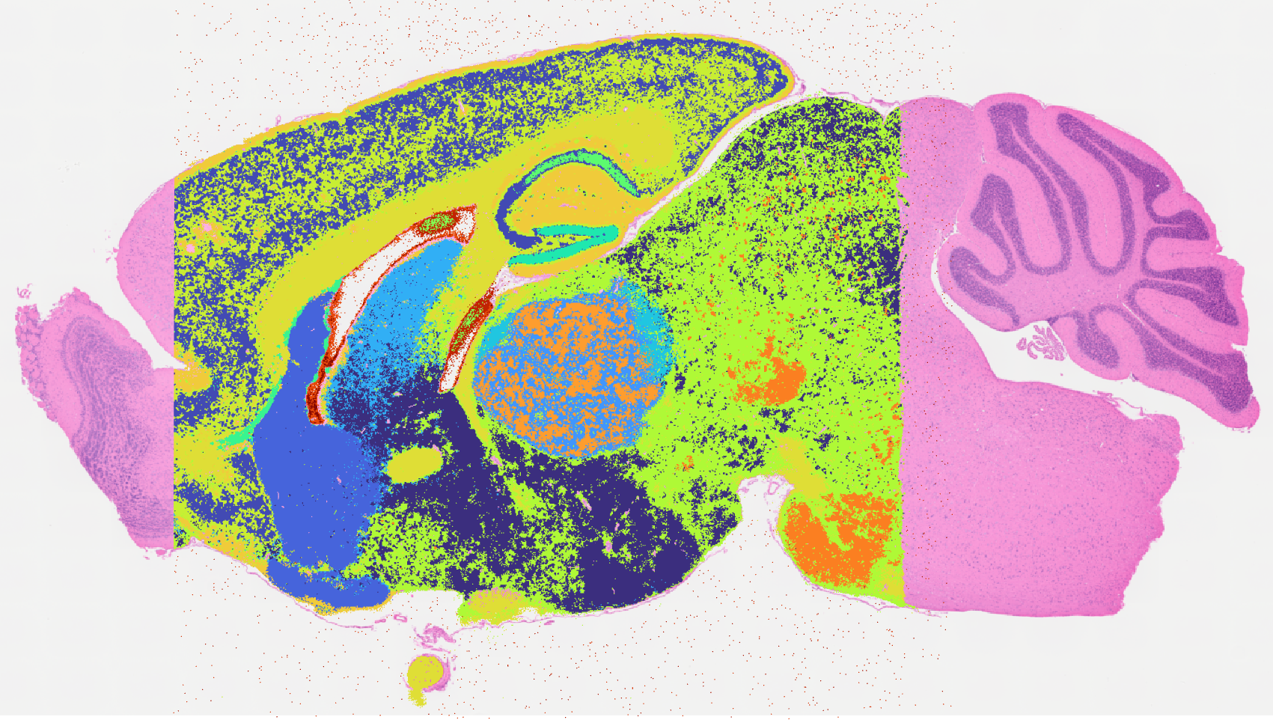
Whole-transcriptome spatial profiling
This pipeline enables spatially resolved transcriptomics across tissue sections, capturing transcriptome-wide gene expression while preserving tissue architecture. It is best suited for:
Unbiased spatial mapping of tissue to discover novel molecular and cellular biomarkers linked to tissue architecture
Revealing spatial patterns of specific biomarkers that arise due disease or in response to treatment
Linking molecular profiles to histology
The workflow integrates tissue preparation, high-resolution imaging, transcriptome capture, sequencing, and downstream analysis into a single coordinated pipeline.
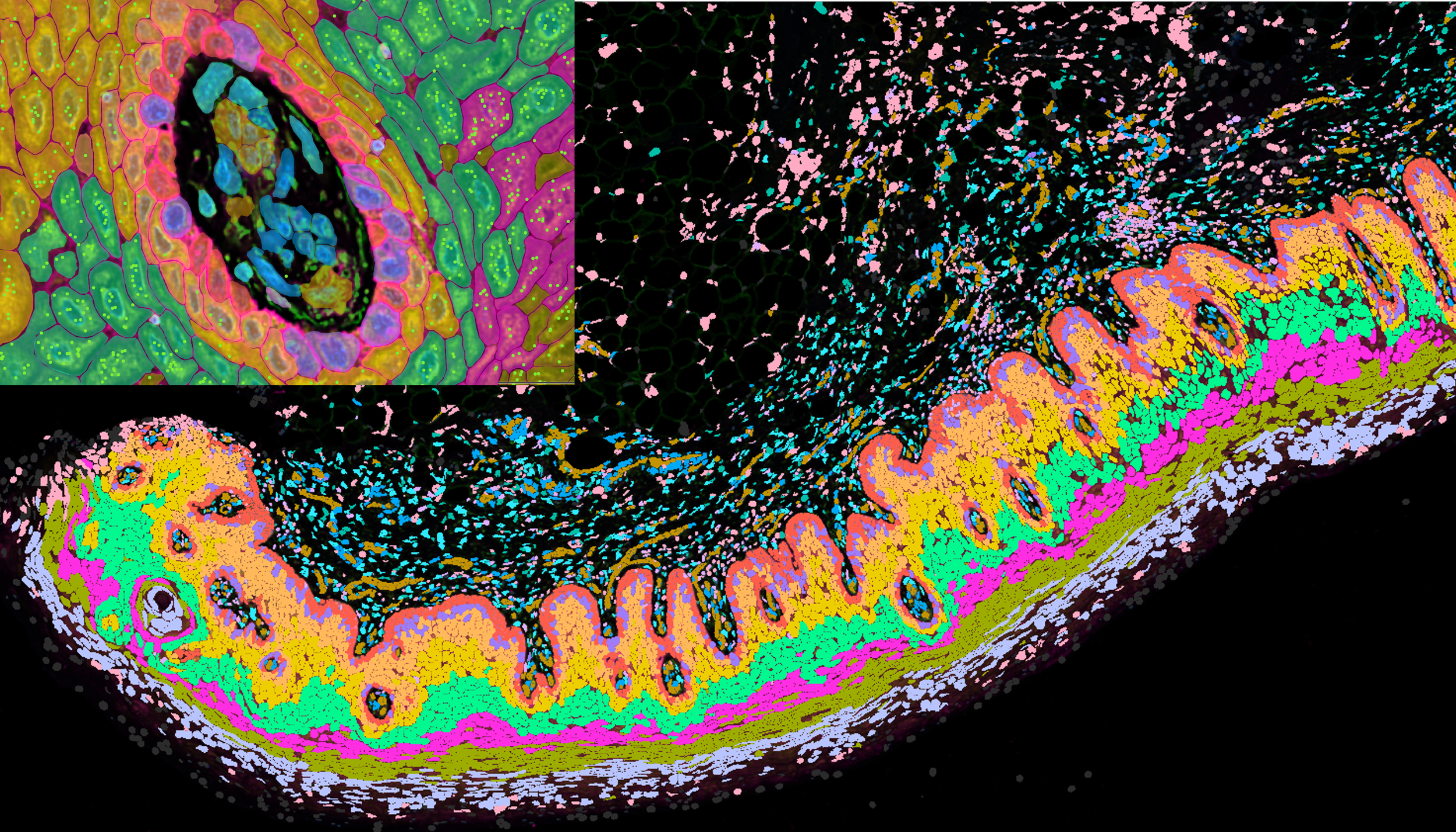
Targeted spatial transcriptomics
This pipeline provides high-resolution, targeted gene expression profiling at subcellular scale, allowing focused interrogation of individual cells and their spatial relationships.
It is particularly valuable when:
Specific gene panels are of interest
Fine spatial resolution is required to characterize or quantify cellular interactions
Analysis needs to be tightly linked to morphology or cell segmentation
Through targeting defined gene sets, this pipeline supports validation of known spatial markers and the analysis of cellular neighbourhoods within tissue microenvironments